Tutorial
This tutorial focuses on using the DeepRetro GUI for retrosynthesis analysis and pathway visualization.
Getting Started
Prerequisites
Run the DeepRetro backend server before opening the GUI.
Keep a valid API key available for frontend access.
Use a modern browser such as Chrome, Firefox, or Safari.
Setup
Open
viewer/index.html.Enter your frontend API key when prompted.
Choose an analysis mode: Smart Retrosynthesis or View Pathway.
Advanced Settings Configuration
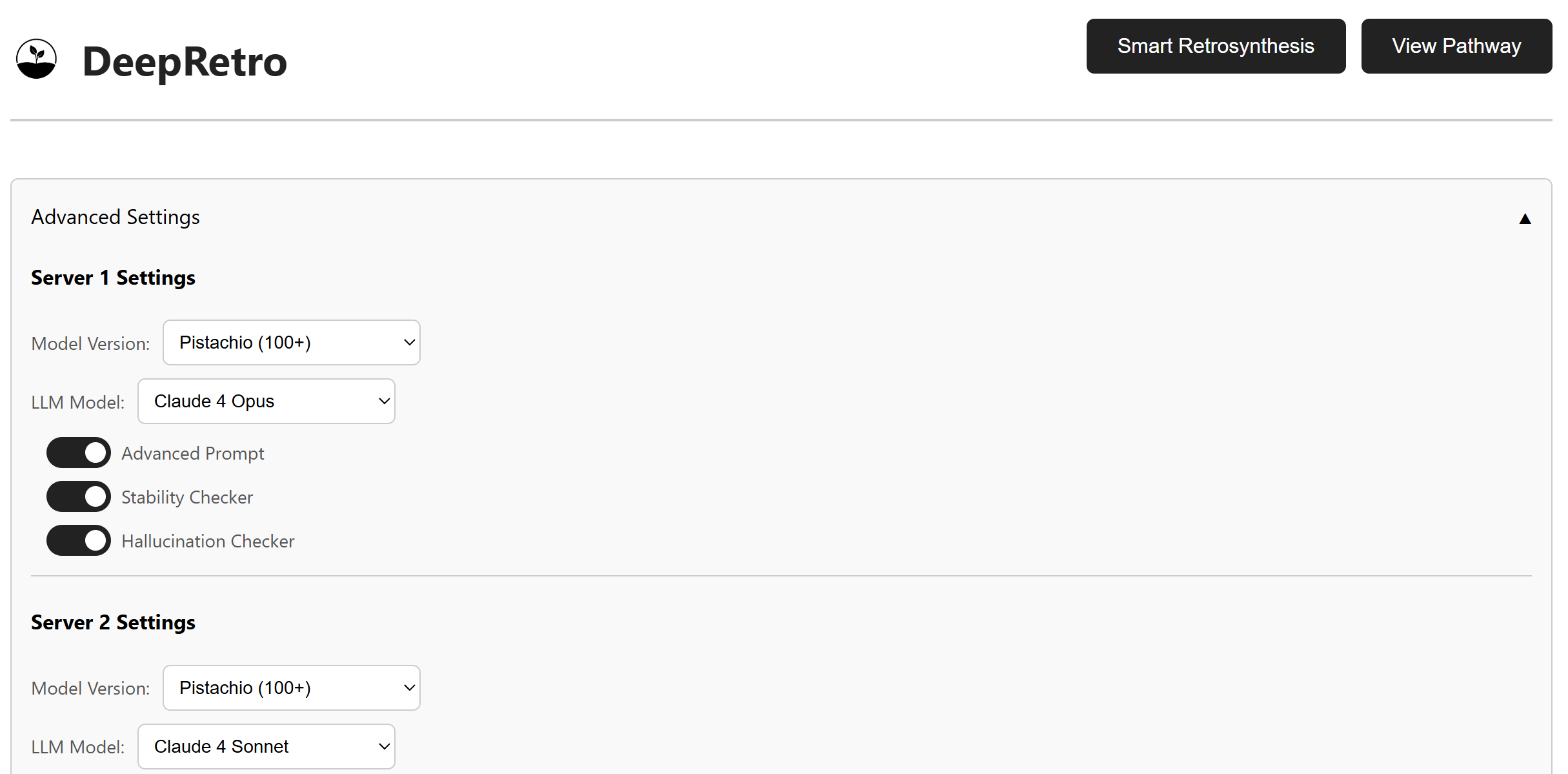
The advanced settings panel exposes the main controls for tuning a run. The default configuration is usually a good starting point, but you can adjust the LLM, backend model, and validation checks to match your use case.
LLM Selection:
Claude 3 Opus, Claude 3.7 Sonnet, DeepSeek, Claude 4 Opus, Claude 4 Sonnet
Model Backend Selection:
USPTO, Pistachio_25, Pistachio_50, Pistachio_100, Pistachio_100+
Validation Checks:
Stability validation enable/disable, hallucination detection settings,
and chemical feasibility assessment
SMILES Input and Configuration
Target Molecule Entry
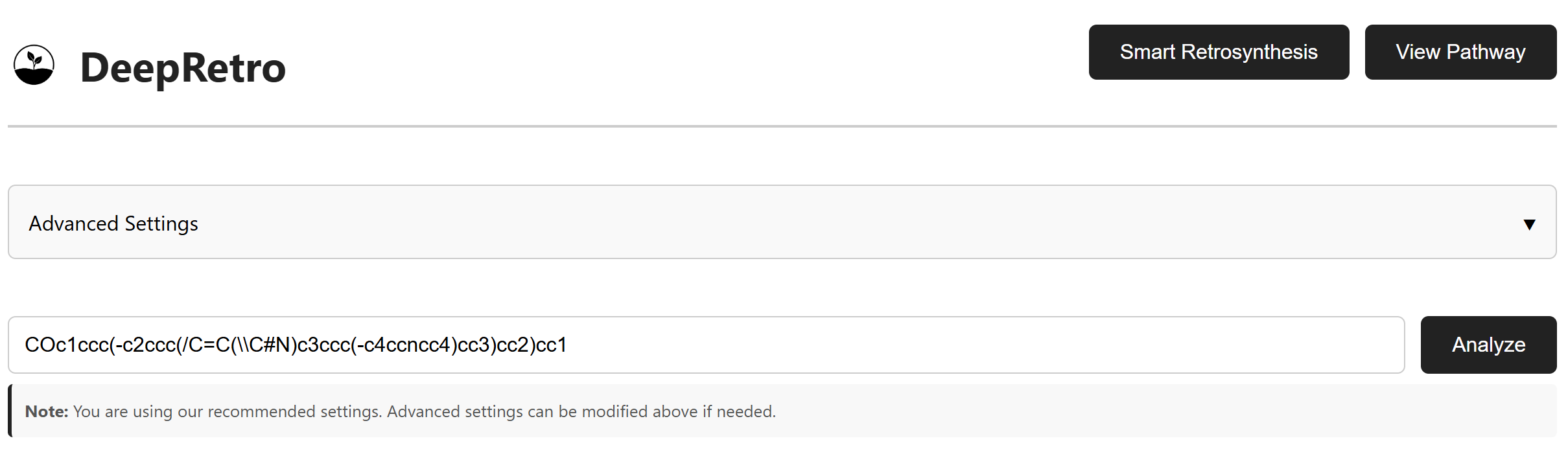
The SMILES input interface allows you to enter the target molecule for retrosynthesis analysis. This tutorial uses Cyanostilbene as an example:
"COc1ccc(-c2ccc(/C=C(\\C#N)c3ccc(-c4ccncc4)cc3)cc2)cc1"
Click Analyze to start the retrosynthesis process. The system validates
the SMILES and begins pathway generation.
Input Requirements
Use valid SMILES notation (Simplified Molecular Input Line Entry System) and choose a chemically feasible target molecule.
Visualization
Pathway Visualization
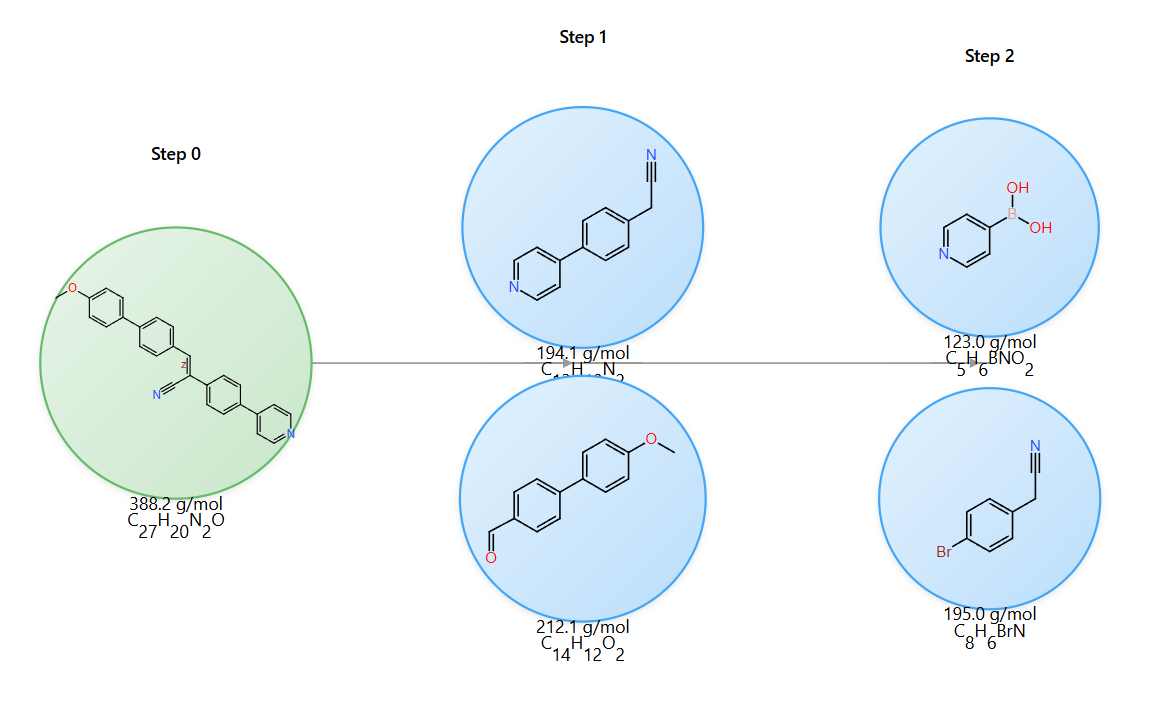
Chemical Context: Cyanostilbene Synthesis
The example demonstrates synthesis of a cyanostilbene derivative, a useful chromophore with aggregation-induced emission and donor-acceptor charge-transfer properties.
Two-step synthesis approach
Step 1 (Knoevenagel condensation): Introduces the cyano group and forms the final conjugated system.
Step 2 (Suzuki coupling): Connects the donor and acceptor aromatic units.
Molecular Information
The visualization provides several interactive elements to help you explore retrosynthesis pathways. Hover on any molecule node to view detailed structural information, including molecular weight, chemical formula, SMILES notation, and step metrics.
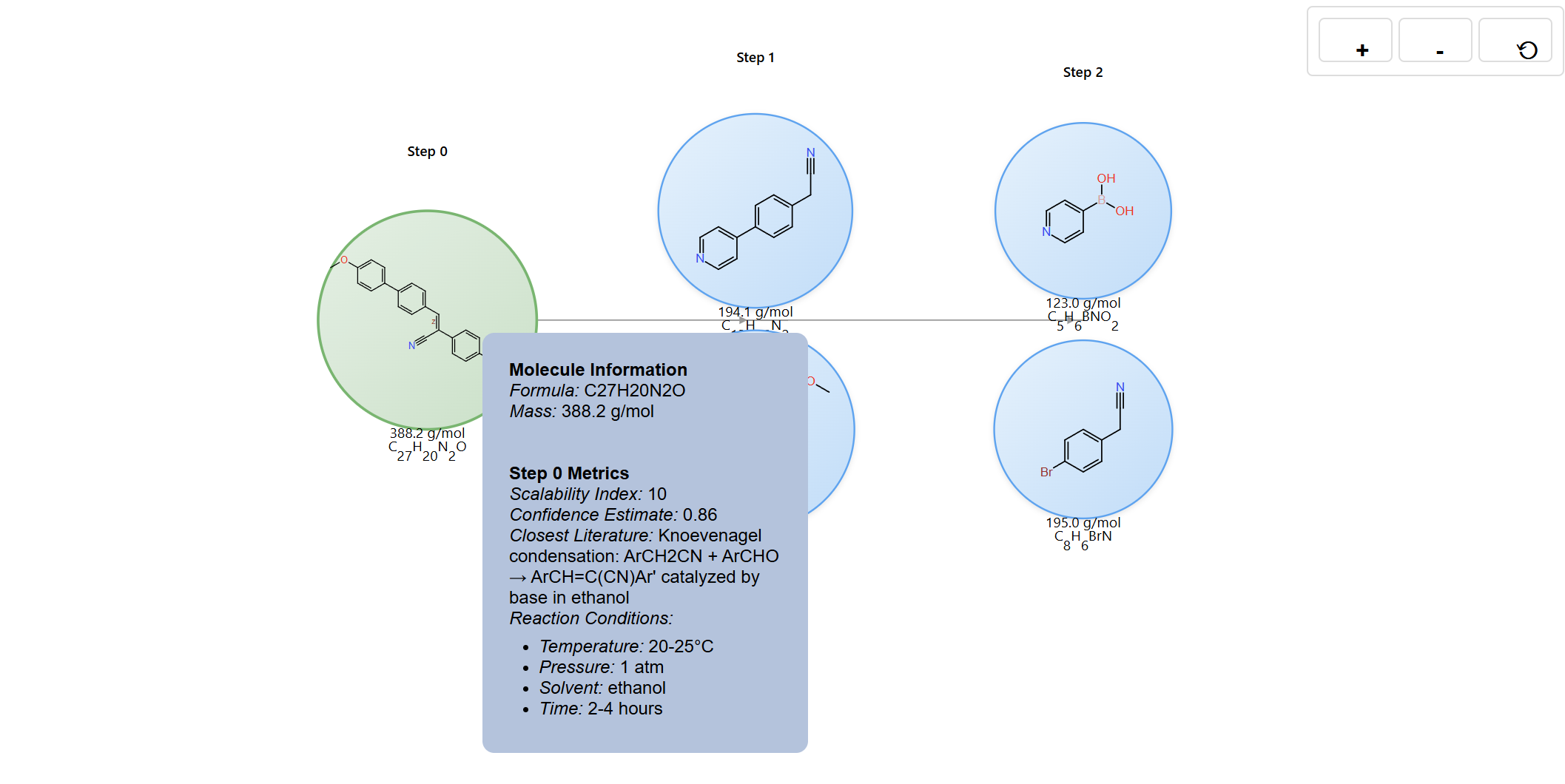
Hover on reaction edges to reveal reaction conditions and metrics, including scalability index, confidence estimate, literature metadata, temperature, and solvent information.
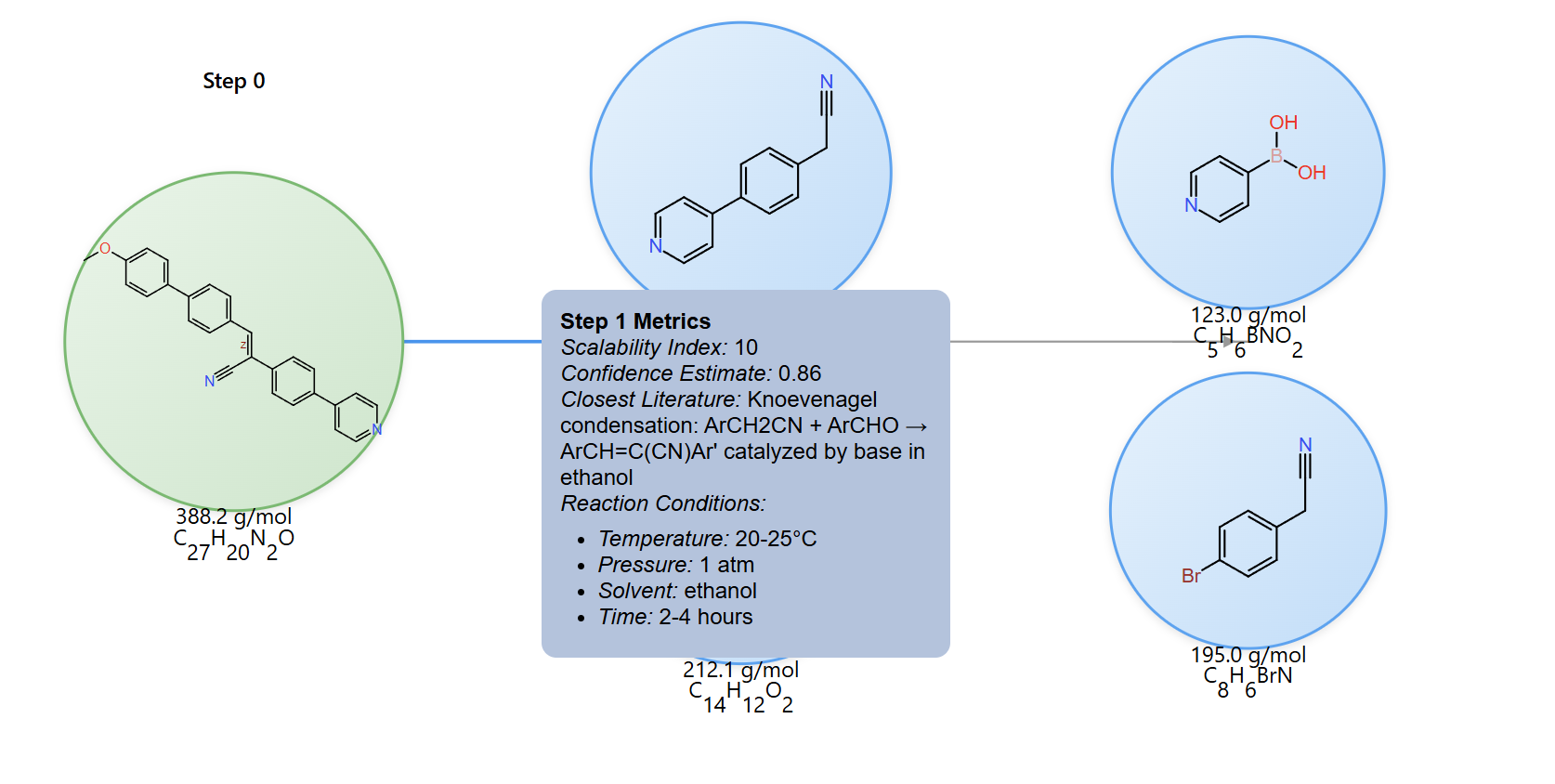
For complex reaction networks, the interface also supports pathway switching, zooming, panning, and step-by-step inspection so you can compare multiple routes side by side.
Interactive Editing
Partial Re-run Analysis
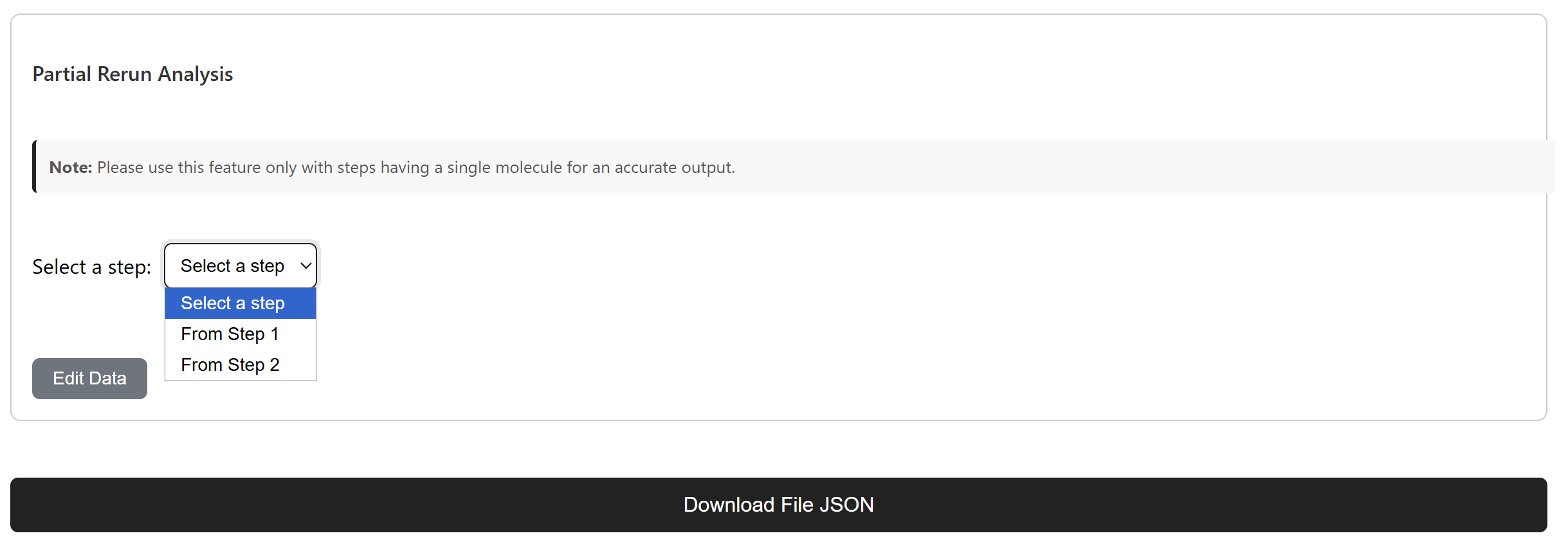
Expert chemist intervention workflow
An expert chemist can select a specific reaction step that needs modification and generate new pathway branches from that point. This supports targeted refinement while preserving successful parts of the route.
Manual Pathway Editing
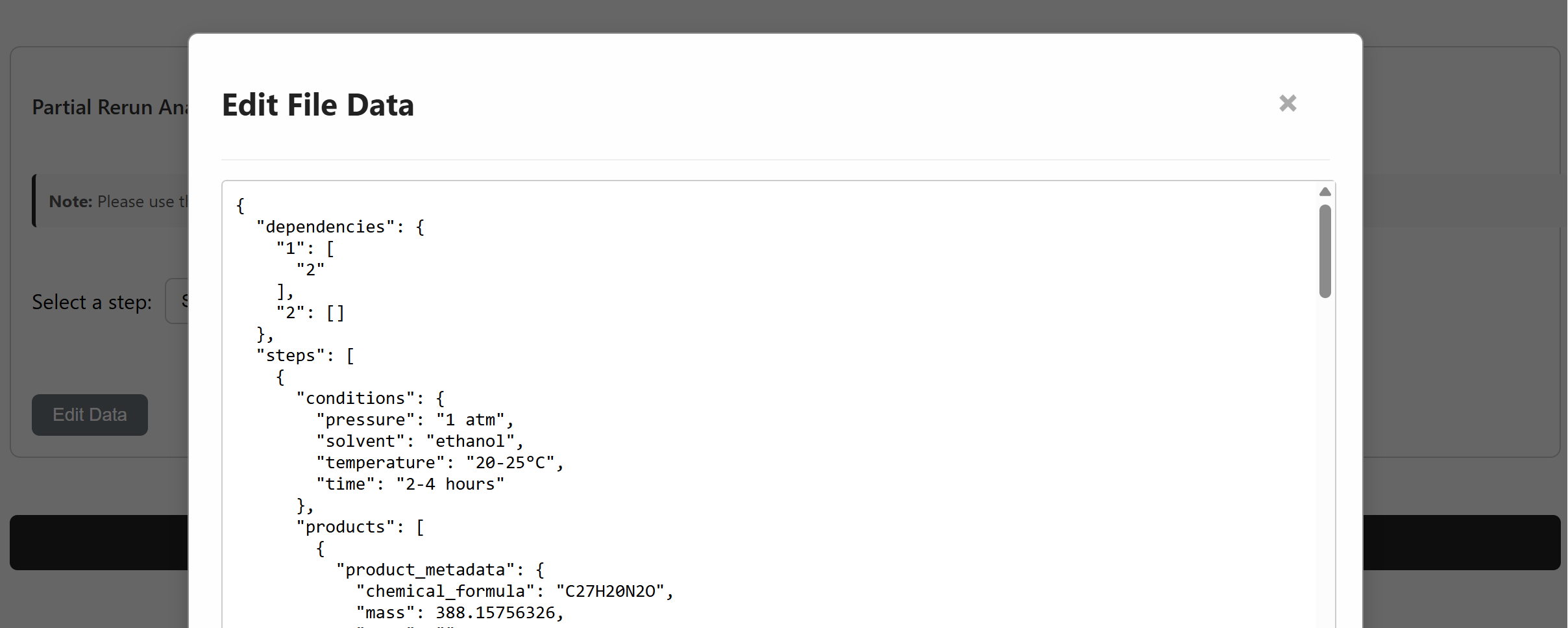
Direct modification
The interface supports manual condition editing, reagent substitution, and
protecting-group addition to refine and optimize reaction pathways. Use
Edit Data to modify the JSON directly, and the pathway view updates
to reflect the changes.
File Management
Pathway File Management
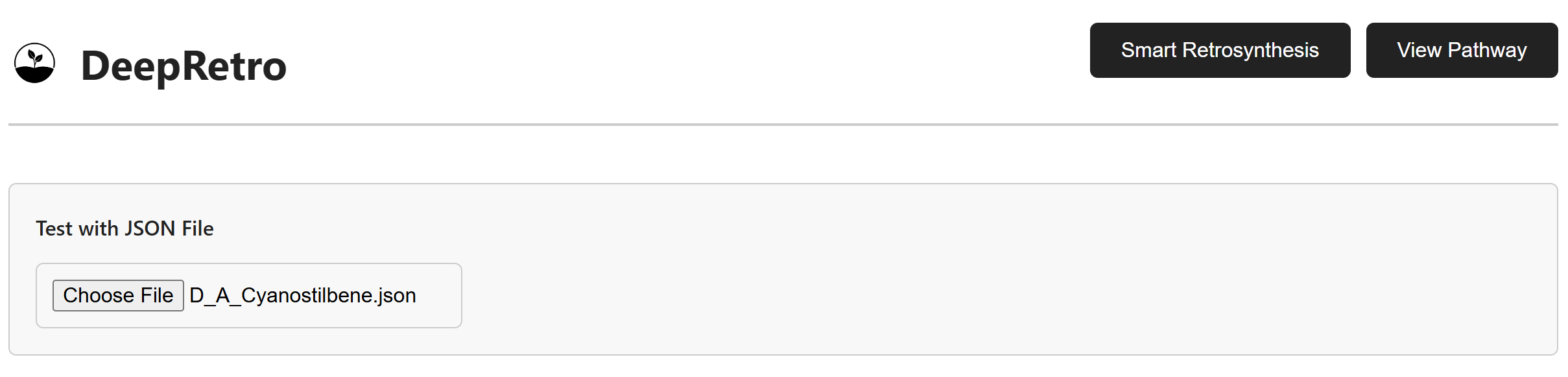
To upload and visualize existing pathways, click the View Pathway tab
and select a JSON file containing pathway data. The system loads and
validates the file before rendering the molecular pathway.
JSON Data Export
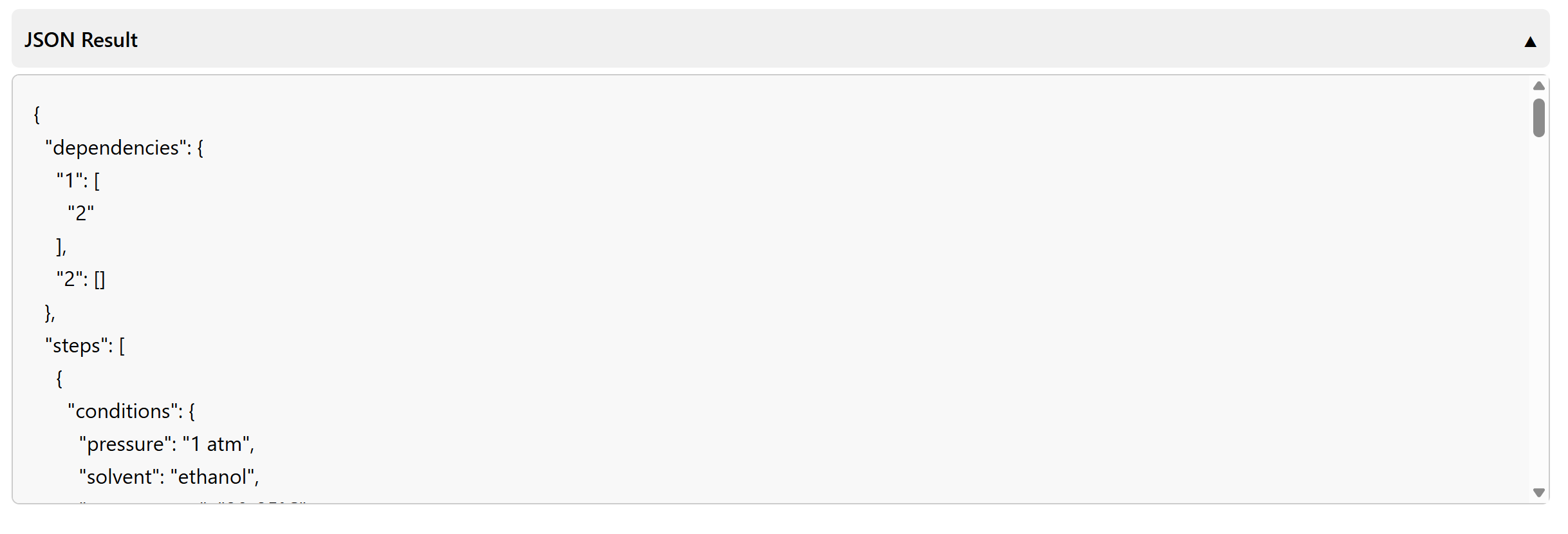
To export pathway data, click JSON Result to inspect the raw JSON and
then use the download control to save it for further analysis or storage.
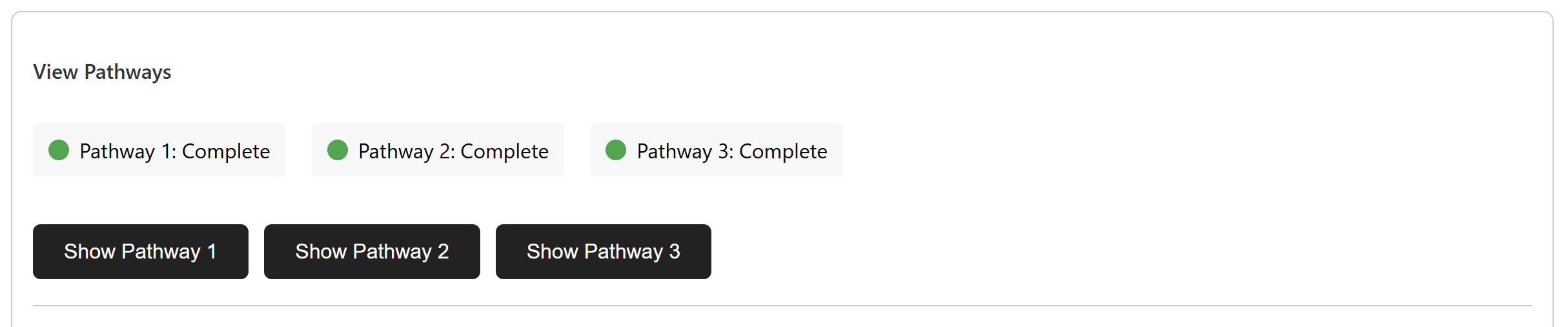
Multiple Pathway Support

The interface supports complex syntheses through pathway switching. You can compare alternative routes, inspect their metrics, and choose the most promising synthetic strategy for the target molecule.
Troubleshooting and Best Practices
Troubleshooting
Common Issues and Solutions
API connection: Verify the server is running, the API key is correct, and the configured URL is reachable.
SMILES errors: Validate the syntax and confirm the target structure is chemically meaningful.
Visualization issues: Refresh the page, clear cached assets if needed, and inspect the browser console for frontend errors.
File upload problems: Confirm the uploaded file is valid JSON and uses the expected pathway schema.
Best Practices
Optimization Guidelines
When working with DeepRetro, start with simple molecules to become familiar with the system. Verify SMILES syntax before submitting a run, enable the relevant validation checks for your use case, and review generated pathways carefully before using them downstream.
For advanced usage, see User Guide, API Reference, and Development Guide.